Can a single experiment do all of the following?
- Provide significant confirmation of the inflationary version of the Big Bang model (and help constrain which model of inflation is correct)
- Confirm the existence of gravitational waves
- Support the quantum nature of gravity (at very high energies)
- Provide the first direct insight into the highest energy levels imagined by physicists – 10^16 GeV (10,000 trillion GeV) – 12 orders of magnitude beyond the LHC
Apparently it can! BICEP2 is a radio telescope experiment located at the South Pole, taking advantage of the very cold, dry air at that remote location for greater sensitivity. It is focused on measuring polarization of the cosmic microwave background radiation that is a remnant of the hot Big Bang of the early universe. (BICEP is an abbreviation of Background Imaging of Cosmic Extragalactic Polarization; this is the second version of the experiment).
The results announced by the BICEP2 team on March 17 at the Harvard-Smithsonian Center for Astrophysics, if they have been correctly interpreted, are the most important in cosmology in the 21st century to date. They are of such enormous significance that a Nobel Prize in Physics is highly likely, if the results and interpretation are confirmed.
We infer from a number of previous observations that there was likely an inflationary period very early on in the universes’s history. We are talking very, very, early – in the first billionth of a trillionth of a trillionth of a second. See this earlier post of mine here: https://darkmatterdarkenergy.com/2011/03/22/inflation/ This new result from BICEP2 is very supportive of inflationary Big Bang models, and that includes very simple models for inflation.
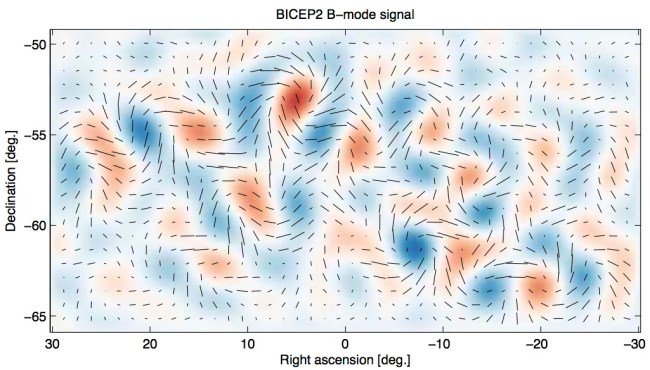
What is the observation? It is B-mode polarization in the cosmic microwave background radiation. The cosmic microwave background (CMB) is the thermal radiation left over from a time when the universe became transparent, at age 380,000 years, almost 14 billion years ago. There are two polarization modes for alignment of CMB radiation, the E-mode and the B-mode. The B-mode measures the amount of “curliness” in the alignment of CMB microwave photons (as you can easily see in the image below).
There are two known causes of B-mode polarization for the CMB. The first, also detected by the BICEP2 experiment, is due to intervening clusters of galaxies along the line of sight. These clusters bend the light paths due to their immense masses, in accordance with general relativity. These effects are seen at smaller spatial scales. At larger spatial scales, we have the more significant effect, whereby the gravitational waves generated during the inflation epoch imprint the polarization.
You can easily see the curly B-mode polarization with a quick glance at BICEP2’s results. (bicepkeck.org)
What is being seen in today’s CMB is due to this second and more profound cause, which is nothing less than quantum fluctuations in space-time in the very, very early universe revealing themselves due to the gravitational waves that they generated. And these gravitational waves in turn caused a small curling effect on the cosmic microwave background, until the time of decoupling of radiation and matter. This is seen in the image at angular scales of a few degrees.
At age 380,000 years the universe became transparent to the CMB radiation and it traveled for another 13.8 billion years and underwent a redshift by a factor of 1500 as the universe expanded. So what was optical radiation at that time, becomes microwave radiation today, with a characteristic temperature of 2.7 Kelvins (degrees above absolute zero), while still retaining the curly pattern seen by BICEP2.
This is the first observation that provides some direct insight into extremely high energy scales within the context of a single experiment. We are talking here of approximately a trillion times higher energy than the Large Hadron Collider, the world’s most powerful particle accelerator, where the Higgs boson was discovered.
The BICEP2 results are a single experiment that for the first time apparently ties quantum mechanics and gravity together. It supports the quantum nature of gravity, which occurs at very high energy scales. The Planck scale at which space-time would be quantized corresponds to an energy level of 10^18 or 10^19 GeV (ten million trillion GeV), and inflation in many models begins when the universe has an energy level somewhat lower, at 10^16 GeV (ten thousand trillion GeV, where 1 GeV is a little more than the rest mass-energy of a proton).
And take a look at this interview of Sean Carroll by PBS’s Gwen Ifill to get some more context around this (hopefully correct!) universe-expanding discovery. Other astronomers are already racing to confirm it.
References:
http://www.cfa.harvard.edu/CMB/bicep2/ – BICEP2 web site at Harvard-Smithsonian Center for Astrophysics
http://www.theguardian.com/science/2014/mar/17/bicep2-how-hot-big-bang-science-dark-energy
http://www.scientificamerican.com/article/gravity-waves-cmb-b-mode-polarization/

April 23rd, 2014 at 11:33 pm
“… there was likely an inflationary period very early on in the universe’s history.” Was this inflation 100% consistent with Newton-Einstein gravitational theory? Or was this inflation consistent with Milgrom’s acceleration law? According to Kroupa, the Lambda-CDM model has been ruled out. On the basis of overwhelming empirical evidence, Milgrom is the Kepler of contemporary cosmology. Is the preceding statement wrong? Note that Kroupa is scheduled to give a talk at the IAS on Thurs., 24 April 2014.
Astrophysics Events, School of Natural Sciences, IAS
September 28th, 2015 at 8:21 am
[…] https://darkmatterdarkenergy.com/2014/03/20/bicep2-apparently-detects-quantum-nature-of-gravity-and-s… […]